Tuan Anh Le
Filtering and smoothing in HMMs and LGSSMs
16 July 2022
A state space model like this
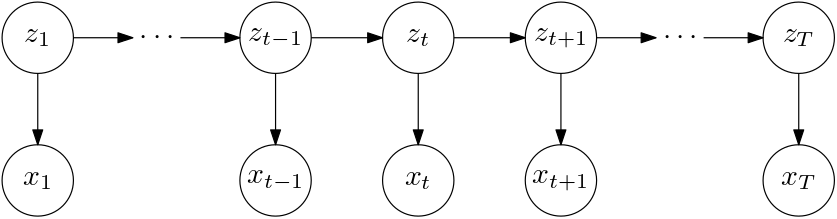
is useful for modelling time series. We’d like to
- Filter: Compute \(p(z_t \given x_{1:t})\) for \(t = 1, \dotsc, T\),
- Smooth: Compute \(p(z_t \given x_{1:T})\) for \(t = 1, \dotsc, T\), and
- Evaluate the full posterior: Sample and compute the probability of \(p(z_{1:T} \given x_{1:T})\). Note that the smoothing distribution above, \(p(z_t \given x_{1:T})\), doesn’t account for the correlations between different latents.
The two most popular instantiations of such a state space model are hidden Markov models (HMMs) and linear Gaussian state space models (LGSSMs). These models are popular because we can filter, smooth and evaluate the full posterior exactly. The algorithms for doing so are called forward-backward for HMMs and Kalman filtering/smoothing for LGSSMs. This note derives those algorithms.
These algorithms are useful building blocks in probabilistic machine learning. They are a special case of belief propagation. There are many good derivation of this algorithm online and in textbooks like Kevin Murphy’s and Christopher Bishop’s. This note provides a derivation that relies on the knowledge of probability rules, conjugate priors and determining whether variables are conditionally independent by using the d-separation rule on a graphical model.
HMMs and LGSSMs
First, let’s define HMMs and LGSSMs.
In HMMs, the initial distribution and transition distributions are categorical distributions
\begin{align}
p(z_1) &= \mathrm{Categorical}(z_1; \pi), \\
p(z_t \given z_{t - 1}) &= \mathrm{Categorical}(z_t; T_{z_{t - 1}}), && (t = 2, \dotsc, T),
\end{align}
where we assume that the latent variables take \(D_z\) values \(z_t \in \{1, \dotsc, D_z\}\); \(\pi\) is a probability vector for \(z_1\) and \(T\) is a matrix where each row \(i\) is a probability vector for transitioning from state \(i\).
For now, let’s assume that the emission / observation distribution is also categorical
\begin{align}
p(x_t \given z_t) &= \mathrm{Categorical}(x_t; E_{z_t}), && (t = 1, \dotsc, T),
\end{align}
where we assume that the observations take \(D_x\) values \(x_t \in \{1, \dotsc, D_x\}\), and that \(E\) is a matrix where each row \(i\) is a probability vector for the observation \(x_t\) given the latent state being \(z_t = i\).
In LGSSMs, we have everything be Gaussian
\begin{align}
p(z_1) &= \mathrm{Normal}(z_1; \mu_1, \Sigma_1), \\
p(z_t \given z_{t - 1}) &= \mathrm{Normal}(z_t; Az_{t - 1} + b; \Sigma_z), && t = 2, \dotsc, T \\
p(x_t \given z_t) &= \mathrm{Normal}(x_t; Cz_t + d; \Sigma_x), && t = 1, \dotsc, T.
\end{align}
The latents and observations can have different dimensions which determines the shapes of the matrix and vector parameters.
Filtering
The goal is to compute \(p(z_t \given x_{1:t})\) for \(t = 1, \dotsc, T\), which we are going to do recursively. First, we are going to compute the filtering distribution for \(t = 1\). Then, we are going to compute the filtering distribution for \(t > 1\), assuming we have already computed the filtering distribution for \(t - 1\).
For \(t = 1\), \begin{align} p(z_1 \given x_1) \propto p(z_1) p(x_1 \given z_1), \end{align} so if the prior \(p(z_1)\) is conjugate for the likelihood \(p(x_1 \given z_1)\), the posterior \(p(z_1 \given x_1)\) will be in the same family of distributions as the prior whose parameters can be computed analytically. This is true for both HMMs and LGSSMs.
For \(t > 1\), assume that we have computed the previous filtering distributions, \(\{p(z_\tau \given x_{1:\tau})\}_{\tau = 1}^{t - 1}\), all of which are in the same family of distributions as the the initial distribution \(p(z_1)\). This is either Categorical for HMMs or Gaussian for LGSSMs.
The first thing to note is that we can apply Bayes rule to express the filtering distribution as a joint distribution over \((z_t, x_t)\) divided by the “marginal likelihood” of \(x_t\), treating \(x_{t - 1}\) as always being conditioned on
\begin{align}
p(z_t \given x_{1:t})
&= p(z_t \given x_t, x_{1:t - 1}) \\
&= \frac{p(z_t, x_t \given x_{1:t - 1})}{p(x_t \given x_{1:t - 1})}.
\end{align}
To make use of conjugacy relationships, we would wish treat the above as computing the conjugate posterior from some conjugate prior-likelihood pair. The most natural thing to try is to decompose the “joint” in the numerator as \begin{align} p(z_t, x_t \given x_{1:t - 1}) = p(z_t \given x_{1:t - 1}) p(x_t \given z_t, x_{1:t - 1}), \end{align} where \(p(z_t \given x_{1:t - 1})\) is the “conjugate prior” and \(p(x_t \given z_t, x_{1:t - 1})\) is the corresponding likelihood. Let’s first take care of the likelihood since that’s easier: we use the d-separation rule to determine from the graphical model above that \begin{align} p(x_t \given z_t, x_{1:t - 1}) = p(x_t \given z_t), \end{align} which means that we can use the likelihood provided in the model (Categorical for HMMs or Gaussian for LGSSMs).
Now, how do we compute the \(p(z_t \given x_{1:t - 1})\) term, which we’d hope to be conjugate to the above likelihood. Let’s try to use the previous filtering distribution, \(p(z_{t - 1}\given x_{1:t - 1})\): \begin{align} p(z_t \given x_{1:t - 1}) = \int p(z_{t - 1}\given x_{1:t - 1}) p(z_t \given z_{t - 1}, x_{1:t - 1}) \,\mathrm dz_{t - 1}. \label{eq:12} \end{align} Like before, we can use the d-separation rule to determine from the graphical model that the second term in the integrand can be written as the transition distribution \begin{align} p(z_t \given z_{t - 1}, x_{1:t - 1}) = p(z_t \given z_{t - 1}). \end{align} Hence, in \eqref{eq:12} we can consider \(p(z_{t - 1}\given x_{1:t - 1})\) to be the conjugate prior to the likelihood \(p(z_t \given z_{t - 1}, x_{1:t - 1})\) and the \(p(z_t \given x_{1:t - 1})\) term to be the “marginal likelihood”. Given a conjugate prior-likelihood pair, we can compute the marginal likelihood exactly.
To summarize,
\begin{align}
p(z_t \given x_{1:t})
&\propto p(z_t, x_t \given x_{1:t - 1}) \\
&= p(z_t \given x_{1:t - 1}) p(x_t \given z_t) \\
&= \left( \int p(z_{t - 1}\given x_{1:t - 1}) p(z_t \given z_{t - 1}) \,\mathrm dz_{t - 1} \right) p(x_t \given z_t)
\end{align}
where computing the integral relies on \(p(z_{1:t - 1} \given x_{1:t - 1})\) being a conjugate prior to the “likelihood” \(p(z_t \given z_{t - 1})\) and computing the filtering distribution relies on the result of the integral being a conjugate prior to the “likelihood” \(p(x_t \given z_t)\).
Smoothing
We want to compute \(p(z_t \given x_{1:T})\) for \(t = 1, \dotsc, T\). Like in filtering, we will do this recursively, but in this case we go from \(t = T\) to \(t = 1\). For \(t = T\), we are already done, as we have \(p(z_T \given x_{1:T})\) from the filtering stage.
For \(t < T\), we assume access to \(\{p(z_\tau \given x_{1:T})\}_{\tau = t + 1}^{T}\).
One straightforward way to get started is to take \(p(z_{t + 1} \given x_{1:T})\), form a joint distribution \(p(z_t, z_{t + 1} \given x_{1:T})\) and marginalize out \(z_{t + 1}\):
\begin{align}
p(z_t \given x_{1:T})
&= \int p(z_t, z_{t + 1} \given x_{1:T}) \,\mathrm dz_{t + 1} \\
&= \int p(z_{t + 1} \given x_{1:T}) p(z_t \given z_{t + 1}, x_{1:T}) \,\mathrm dz_{t + 1}.
\end{align}
Since we already have access to \(p(z_{t + 1} \given x_{1:T})\), we only need to worry about \(p(z_t \given z_{t + 1}, x_{1:T})\). The first thing to note is that due to the factorization in the graphical model above, we can drop the dependency on \(x_{t + 1:T}\): \begin{align} p(z_t \given z_{t + 1}, x_{1:T}) &= p(z_t \given z_{t + 1}, x_{1:t}). \end{align}
Finally, let’s treat this as a posterior of \(z_t\) given \(z_{t + 1}\) and write it in terms of a prior and a likelihood:
\begin{align}
p(z_t \given z_{t + 1}, x_{1:t})
&= \frac{p(z_t, z_{t + 1} \given x_{1:t})}{p(z_{t + 1} \given x_{1:t})} \\
&= \frac{p(z_t \given x_{1:t}) p(z_{t + 1} \given z_t, x_{1:t})}{p(z_{t + 1} \given x_{1:t})}
\end{align}
where the \(p(z_t \given x_{1:t})\) is the “prior” and \(p(z_{t + 1} \given z_t, x_{1:t})\) is the “likelihood” which can be simplified to \(p(z_{t + 1} \given z_t)\) thanks to the factorization of the generative model.
We have access to \(p(z_t \given x_{1:t})\) from filtering and \(p(z_{t + 1} \given z_t)\) is the transition distribution.
The “prior” is conjugate to the “likelihood”, hence we can compute \(p(z_t \given z_{t + 1}, x_{1:T})\) exactly.
To summarize,
\begin{align}
p(z_t \given x_{1:T})
&= \int p(z_t, z_{t + 1} \given x_{1:T}) \,\mathrm dz_{t + 1} \\
&= \int p(z_{t + 1} \given x_{1:T}) p(z_t \given z_{t + 1}, x_{1:T}) \,\mathrm dz_{t + 1} \\
&= \int p(z_{t + 1} \given x_{1:T}) p(z_t \given z_{t + 1}, x_{1:t}) \,\mathrm dz_{t + 1} \\
&= \int p(z_{t + 1} \given x_{1:T}) \frac{p(z_t \given x_{1:t}) p(z_{t + 1} \given z_t, x_{1:t})}{p(z_{t + 1} \given x_{1:t})} \,\mathrm dz_{t + 1} \\
&= \int p(z_{t + 1} \given x_{1:T}) \frac{p(z_t \given x_{1:t}) p(z_{t + 1} \given z_t)}{p(z_{t + 1} \given x_{1:t})} \,\mathrm dz_{t + 1},
\end{align}
where the fraction in the integrand equals the conjugate posterior for the prior \(p(z_t \given x_{1:t})\) and likelihood \(p(z_{t + 1} \given z_t)\).
This conjugate prior is then used as the “likelihood” for the conjugate prior \(p(z_{t + 1} \given x_{1:T})\) and the result of integration is the “marginal likelihood”.
Evaluating the full the posterior
Here, we’d like to evaluate the probability of the full posterior \(p(z_{1:T} \given x_{1:T})\) and sample from it given a sequence of observations \(x_{1:T}\).
From the previous section, we know that we can access \(p(z_T \given x_{1:T})\) as well as
\begin{align}
p(z_t \given z_{t + 1:T}, x_{1:T})
&= p(z_t \given z_{t + 1}, x_{1:T}) \\
&\propto p(z_t \given x_{1:t}) p(z_{t + 1} \given z_t)
\end{align}
for \(t < T\) where the first equality holds due to \(z_t\) being independent of \(z_{t + 2:T}\) given \(z_{t + 1}\).
This can be computed as a conjugate posterior for the prior \(p(z_t \given x_{1:t})\) (obtained from the filtering step) and likelihood \(p(z_{t + 1} \given z_t)\) (which is the transition distribution of the model).
Hence, we can write \begin{align} p(z_{1:T} \given x_{1:T}) = p(z_T \given x_{1:T}) \prod_{t = T - 1, \dotsc, 1} p(z_t \given z_{t + 1:T}, x_{1:T}) \end{align} which we can sample from ancestrally and evaluate the probability by plugging in \(z_{1:T}\) directly.
[back]